There is a simple setup dialog that lets you control basic search characteristics, including choosing the specific type of nuclease to be used (we currently include 32 different characterized motifs in the default PAM file). One new function introduced a few years ago will search for PAM sequences in your sequence. The most commonly used Cas9 nuclease (SpCas9) from Streptococcus pyogenes recognizes the PAM sequence NGG. Once activated, your key will enable MacVector to submit up to 10 requests per second irrespective of other users at your Institution, which will dramatically speed up population of the BLAST Map tab and other NCBI services, particularly in shared environments.ĬRISPR-Cas9 genetic editing mechanisms require a short (typically 20nt) RNA sequence complementary to a target site next to a Protospacer Adjacent Motif (PAM) sequence. Once you have a key, you can register it in MacVector using the MacVector | Preferences | Internet tab and pasting your API key into the Entrez Server edit box You can get around this by registering for your own personal NCBI API key. While most users may not see a major difference with this, if you are sharing an institutional IP address with other MacVector users, you may find that the BLAST Map tab is slow to populate, and other NCBI-related functions may also be slow. This can really help you see the context of your hits, and let you download just the relevant region for additional analysis.Ī few years ago the NCBI introduced a “throttling” procedure such that unregistered users are limited to three requests per second per IP address. 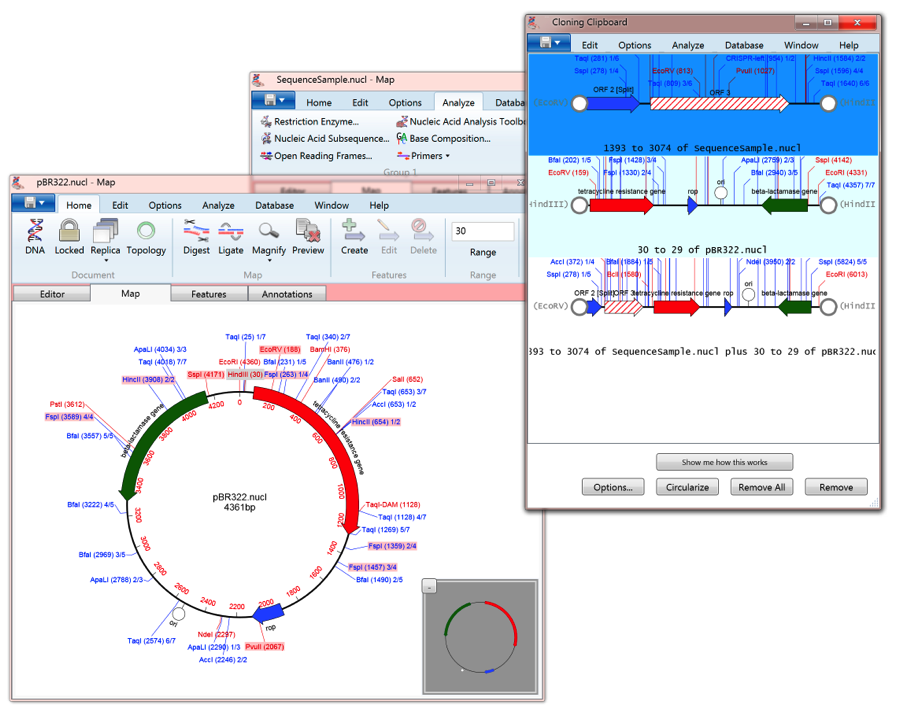
Particularly if you are working with prokaryotic sequences, where most BLAST hits these days are to genome sequences. MacVector has a very cool BLAST Map results tab that displays the annotations surrounding any hits from the selected database. and don’t forget that if you prefer Ligase Independent Cloning or other similar techniques the tool will do those too. We do not currently have a dedicated tutorial for Gibson/Ligase-independent Assembly, but there is a useful section with examples you can follow in the Whats New in MacVector Workshop manual. (Note the gray CC residues that were inserted to ensure that the adjacent ATG start codon is maintained in-frame) In addition, the interface lets you view the translations around the junctions so you can be absolutely sure that your primer will create that perfect fusion protein. The algorithm is very similar, but you can add your annotated MacVector nucleic acid files and the final molecule will retain all of their features and annotations. If you are familiar with the New England Biolabs NEBuilder interface then you will love this. From there, you can choose the type of assembly (it doesn’t have to be the usual 5’ exonuclease Gibson approach) and follow the instructions. All you have to do to get started is choose the File->New->Gibson/Ligase-independent Assembly… menu item. You can use MacVector to design primers for multi-fragment Gibson Assembly, and also generate the predicted recombinant DNA molecule resulting from the assembly.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
